Ladurner Lab
Molecular mechanisms of chromatin plasticity
Molecular mechanisms of chromatin plasticity

© J. Greune
Our team is interested in identifying, characterizing and then exploiting novel molecular mechanisms that underlie this level of regulatory plasticity in chromatin structure. We are uncovering fundamental mechanisms that govern the function of our complex genome and its ability to adapt to new environments. We are studying the role of post-translational modifications and of cellular metabolites in regulating the assembly and remodelling of chromatin.
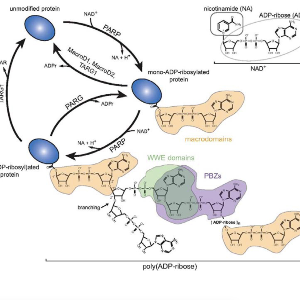
DNA damage is a major driver of genomic instability, cancer initiation, and therapy resistance, making its regulation central to human disease. Our research focuses on defining the biological functions and cell biology of ADP-ribosylation in the DNA damage response. We investigate how the DNA damage–activated ADP-ribosyltransferase PARP1 is regulated through recognition of DNA strand breaks, and how its enzymatic activation drives nucleosome remodeling by the molecular machine ALC1 (CHD1L). ALC1 plays a critical role in cancer biology, and our work has enabled the development of an allosteric small-molecule inhibitor that is currently in clinical trials.
Dietary carbohydrates such as glucose and fructose play central roles in metabolic homeostasis but, when in excess, contribute to metabolic syndrome by activating gene programs that promote lipid synthesis. Our research addresses the fundamental question of how cellular metabolites directly regulate gene activity to mediate adaptive and potentially also pathophysiological responses to nutritional changes. We discovered the first cellular receptor for a key glucose-derived metabolite, demonstrating that this endogenous metabolite and its chemical analogs function as high-affinity molecular switches (agonists and antagonists) for a conserved human transcription factor. Building on this work, we are dissecting the molecular mechanisms by which metabolites regulate transcription factor activity and developing synthetic small molecules to modulate these interactions. This research has strong translational potential for the treatment of metabolic syndromes and is also highly relevant to cancer, where related metabolite–protein interactions drive aberrant gene regulation and can be therapeutically targeted.
| Name | Position | |
|---|---|---|
| Heimhalt, Maren | maren.heimhalt | Postdoc |
| Kotthoff, Christiane | christiane.kotthoff | Research Technician |
| Ladurner, Andreas | andreas.ladurner | Chair of the Department; Group Leader |
| Lee, Hannah | hannah.lee | PhD Student |
| Nagel, Hanna | hanna.nagel | PhD Student |
| Preißer, Julia | julia.preisser | Research Technician |
| Subramanian, Swathi | swathi.subramanian | PhD Student |
| Usha, Sandra | sandra.usha | Postdoc |
| Wright, Tanner | tanner.wright | Postdoc |