Colomé-Tatché Lab
Computational Epigenetics
Computational Epigenetics
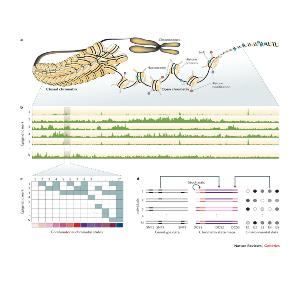
We are interested in understanding how different epigenomes emerge, how stable epigenetic changes are, and how they lead to different phenotypes. In order to answer these questions, we develop computational methods and mathematical models to determine what regions of the epigenome are altered under different conditions in large numbers of individuals or in single cells, and how these changes affect the observed phenotype.
We work on the development of computational models for the analysis of single cell DNA methylation data (i.e. single cell bisulfite sequencing) and single cell open chromatin data (i.e. scATAC-seq), and integration with single cell transcriptomics data. We have developed an R package for the calling of CNVs from single cell genomics data (aneufinder (https://bioconductor.org/packages/release/bioc/html/AneuFinder.html)) and a python package for the analysis of single cell DNA methylation data and single cell open chromatin data (performing common clustering, dimension reduction, trajectory learning, data integration and differential calling) (epiScanpy (https://episcanpy.readthedocs.io/en/latest/)).
We work on the development of improved analysis tools for the integration of several epigenetic marks, like DNA methylation and several histone modifications or chromatin openness. We are mainly interested in integrating large number of measurements in the same sample, or in situations where an epigenetic mark has been measured in a large number of samples or time points.
We are interested in the modelling of dynamic epigenetic processes, such as DNA methylation inheritance. We are interested in epigenetic inheritance in plants, where it is well established that changes in DNA methylation can be transmitted across generations via meiotic inheritance.
We work on different biological and medical applications, in strong collaboration with experimental partners. Our goal is to apply our developed methods to the analysis and interpretation of different sorts of single-cell and/or bulk multi-omic data.
| Name | Position | |
|---|---|---|
| Celik, Muhammet | muhammet.celik | PhD Student |
| Colomé-Tatché, Maria | maria.colomé-tatché | Assistant Professor of Functional Genomics and Cell Biology |
| Malagoli, Gabriele | gabriele.malagoli | PhD Student |
| Schmid, Katharina | katharina.schmid | Postdoc |
| Symeonidi, Aikaterina | katia.symeonidi | Postdoc |
| Tueysuez, Irem | irem.tueysuez | PhD Student |