Mock Lab
Max-Eder Research Group - Clinical Multi-Omics
Max-Eder Research Group - Clinical Multi-Omics
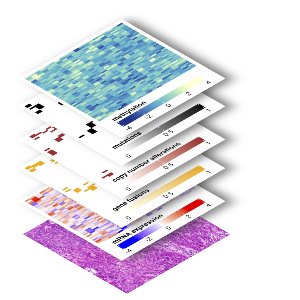

© Andreas Mock
Cancer is a complex and heterogeneous disease that cannot be captured by a single data type. Clinical multi-omics addresses this by integrating diverse molecular layers—including genomics, epigenomics, transcriptomics, proteomics, and metabolomics—into a unified view of tumor biology.
This framework is further strengthened by single-cell technologies, spatial profiling, and AI-driven digital pathology, which add resolution and functional context. Single-cell approaches resolve tumor heterogeneity at the level of individual cells, uncovering rare populations and dynamic cellular states.
Spatial profiling places these molecular features into their anatomical context, enabling the analysis of tumor–microenvironment interactions, including immune and stromal compartments. At the same time, digital pathology provides quantitative, high-throughput analysis of tissue morphology and links histological patterns to molecular alterations.
Together, these approaches enable a comprehensive and spatially resolved understanding of cancer, forming the basis for more precise diagnostics and targeted therapeutic strategies.
HE staining serve as training data for predictions and performance can be e.g. tested by (multiplex) immunohistochemistry. | © Andreas Mock
Digital pathology can combine high-resolution histology with computational analysis to quantitatively link tissue morphology to molecular states. AI now enables the inference of gene expression directly from routine H&E sections.
Recent advances, such as the generative AI framework Phoenix, establish virtual spatial transcriptomics, predicting spatially resolved gene expression from standard histology and enabling large-scale analysis of archived pathology material.
This approach enables the reconstruction of tissue ecosystems, capturing cell states, spatial niches, and cell–cell interactions, and supports in silico modeling of treatment response.
By overcoming the cost and scalability limitations of experimental spatial methods, virtual spatial transcriptomics provides a framework for population-scale morphomolecular profiling in precision oncology.
We are part of the Munich Center for Computational Pathology (MCCP).
Pan-cancer virtual spatial transcriptomics from routine histology with Phoenix
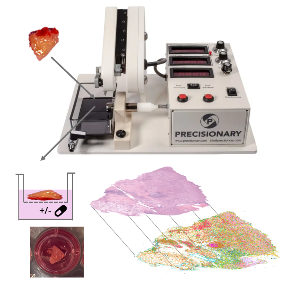

Fresh tumor tissue is obtained from the operating room and processed by a vibrating microtome and cultivated at the air-liquid-interface. Deep biological readouts can be achied by spatial profiling methods. | © Andreas Mock
Organotypic tumor slice cultures preserve the native architecture, cellular heterogeneity, and microenvironment of patient-derived tissues, providing a powerful platform for functional precision oncology. Fresh tumor specimens are processed into viable slices and cultivated at the air–liquid interface, allowing therapies to be tested directly in the patient’s own tissue context.
Combined with single-cell and spatial profiling, this approach enables spatially resolved ex vivo drug response analysis, linking therapeutic effects to defined tumor cell states and microenvironmental interactions. Our recent study on SMARCB1-deficient sinonasal carcinoma illustrates how this strategy supports rare cancer research, where established preclinical models are lacking, by identifying distinct malignant subpopulations and revealing a selective therapeutic vulnerability to mTOR inhibition.
Tumor slice cultures bridge experimental modeling and clinical translation, enabling mechanism-informed therapy testing directly from patient material.
Spatially resolved ex vivo drug response profiling in SMARCB1-deficient sinonasal carcinoma
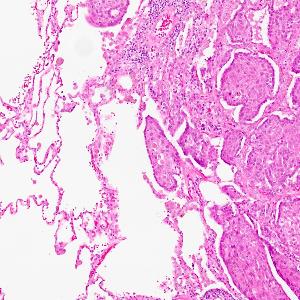
Head and neck cancer cells (right) infiltrate into the lung (left). | © Andreas Mock
Head and neck squamous cell carcinoma (HNSCC) is the seventh most common cancer worldwide, accounting for ~4.5% of cancer-related deaths. More than 60% of patients present with locally advanced disease (stage III/IV), associated with a high risk of recurrence and distant metastasis.
Despite advances in multimodal therapy (surgery, radiotherapy, and systemic treatments), outcomes for patients with recurrent or metastatic disease remain poor, highlighting a critical need for new therapeutic strategies.
By integrating single-cell sequencing, tumor slice cultures, spatial tumor profiling, and multimodal data analysis, we aim to identify mechanism-based vulnerabilities and develop novel therapeutic approaches for incurable HNSCC. This integrative framework enables the study of tumor heterogeneity, microenvironmental interactions, and treatment response directly in patient-derived material.
Our head and neck cancer research is supported by the Max-Eder Junior Research Group Program of the German Cancer Aid (Deutsche Krebshilfe)

Assistenzarzt & Arbeitsgruppenleiter
Max-Eder Research Group - Clinical Multi-Omics
Senior Bioinformatician
Clinical Bioinformatics
Research Associate
Spatially resolved ex vivo drug response profiling in SMARCB1-deficient sinonasal carcinoma
Jurmeister P*, Flach S*, Bergmayr L, Schleich K, Calderon EC, Mochmann LH, Zhdanovich Y, Klingler D, Pusztai A, Kübler A, Walz C, Westphalen CB, Beck A, Leitheiser M, Breimer GE, Rijken JA, Devriese L, Baumeister P, Skálová A, Schallenberg S, Klauschen F*, Mock A*.
EMBO Mol Med. 2026 doi.org/10.1038/s44321-026-00437-1
Pan-cancer virtual spatial transcriptomics from routine histology with Phoenix
Tran M*, Gindra RH*, Putze P, Senbai K, Palla G, Kos T, Falcomatà C, Wang C, Guo R, Boxberg M, Berclaz LM, Lindner LH, Bergmayr L, Knösel T, Jurmeister P, Klauschen F, Homicsko K, Gottardo R, Eckstein M, Matek C, Mock A*, Theis FJ*, Sauer D*, Peng T*.
bioRxiv. 2026 doi: doi.org/10.64898/2026.04.25.720812
Beyond Attention Heatmaps: How to Get Better Explanations for Multiple Instance Learning Models in Histopathology
Jamshidi Idaji M*, Hense J*, Neuhäuser T, Krause A, Luo Y, Eberle O, Schnake T, Ciernik L, Rezaei Jafari F, Vahidimajd R, Dippel J, Walz C, Klauschen F, Mock A, Müller K-R.
arXiv. 2026 doi.org/10.48550/arXiv.2603.08328
Digital Spatial Pathway Mapping Reveals Prognostic Tumor States in Head and Neck Cancer
Hense J*, Jamshidi Idaji M*, Ciernik L, Dippel J, Ersan F, Knebel M, Pusztai A, Sendelhofert A, Buchstab O, Fröhling S, Otto S, Hess J, Liokatis P, Klauschen F, Müller K-R, Mock A
bioRxiv. 2025 doi: 10.1101/2025.11.24.689710
EGFR-mediated local invasiveness and response to Cetuximab in head and neck cancer
Zhou J, He M, Zhao Q, Shi E, Wang H, Ponksche V, Song J, Wu Z, Ji D, Kranz G, Tscherne A, Schwenk-Zieger S, Razak NA, Hess J, Belka C, Zitzelsberger H, Ourailidis I, Stögbauer F, Boxberg M, Budczies J, Reichel CA, Canis M, Baumeister P, Wang H, Unger K, Mock A, Gires O
Mol Cancer. 2025 Mar 22;24(1):94. doi: 10.1186/s12943-025-02290-1
NCT/DKFZ MASTER handbook of interpreting whole-genome, transcriptome, and methylome data for precision oncology
Mock A*, Teleanu MV*, Kreutzfeldt S, Heilig CE, Hüllein J, Möhrmann L, Jahn A, Hanf D, Kerle IA, Singh HM, Hutter B, Uhrig S, Fröhlich M, Neumann O, Hartig A, Brückmann S, Hirsch S, Grund K, Dikow N, Lipka DB, Renner M, Bhatti IA, Apostolidis L, Schlenk RF, Schaaf CP, Stenzinger A, Schröck E, Hübschmann D, Heining C, Horak P, Glimm H, Fröhling S.
NPJ Precis Oncol. 2023 Oct 26;7(1):109. doi: 10.1038/s41698-023-00458-w
Transcriptome profiling for precision cancer medicine using shallow nanopore cDNA sequencing
Mock A, Braun M, Scholl C, Fröhling S, Erkut C.
Sci Rep. 2023 Feb 9;13(1):2378. doi: 10.1038/s41598-023-29550-8
Comprehensive Genomic and Epigenomic Analysis in Cancer of Unknown Primary Guides Molecularly-Informed Therapies Despite Heterogeneity
Möhrmann L*, Werner M*, Oleś M*, Mock A*, Uhrig S, Jahn A, Kreutzfeldt S, Fröhlich M, Hutter B, Paramasivam N, Richter D, Beck K, Winter U, Pfütze K, Heilig CE, Teleanu V, Lipka DB, Zapatka M, Hanf D, List C, Allgäuer M, Penzel R, Rüter G, Jelas I, Hamacher R, Falkenhorst J, Wagner S, Brandts CH, Boerries M, Illert AL, Metzeler KH, Westphalen CB, Desuki A, Kindler T, Folprecht G, Weichert W, Brors B, Stenzinger A, Schröck E, Hübschmann D, Horak P, Heining C, Fröhling S, Glimm H.
Nat Commun. 2022 Aug 2;13(1):4485. doi: 10.1038/s41467-022-31866-4